Signal-passing Tile Assembly Model (STAM)
The signal tile assembly model(STAM) describes a signaled tile self assembly process that enriches the tile assembly paradigm with improved capabilities and allows tile assembly to more closely emulate biological and natural processes.
Signaled glue activation was introduced for the purpose of allowing supertiles to take on new identities or roles once assembled [2], so that supertile interactions as described in hierarchical models such as the Two-Handed Assembly Model (2HAM) can simulate the interactions of individual tiles.
The generalized model presented here has been designed to take into consideration a DNA implementation of all aspects of signaled assembly: binding, signaling, and glue activation or deactivation.
Physical Basis for the Model
The physical body of a tile might be composed entirely from a DNA origami structure which has more room for signal pathways than smaller DNA structures that have been used to implement the passive tiles of the TAM. Toe-hold mediated DNA strand exchange mechanisms [3-6], as shown in Figure 1, provide a basis for a plausible physical implementation of signaling, glue activation and glue deactivation. Signals across individual tiles could potentially be implemented using DNA hybridization cascades such as HCR [7], transducers [8], seesaw gates [9], other mechanisms [10], or by DNA walkers [11-14].
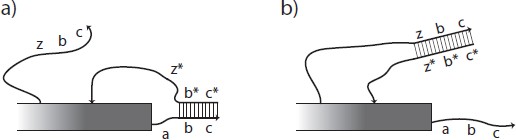
Fig. 1: Activation of a latent glue. Single-stranded DNA is depicted as a line with an arrow at the 3' end and double- stranded DNA is shown as two parallel lines connected by a ladder of base pairs. Letters such as a and y indicate short sequences of DNA bases and a star indicates the complementary sequence. Rectangles indicate structures, possibly composed of DNA origami, to which the relevant sticky ends of DNA are attached. a) The strand zbc is presumed to be unavailable until freed by a previous signaling cascade (not shown). The sequence labeled z may then bind at the toehold z8 on the protection strand c* b* z*, removing it from the glue strand abc via strand displacement. The protection strand is shown bound only to b and c, which is enough to prevent interaction with the complementary tile edge shown as long as no toeholds are free to interact. Alternatively, the entire sequence abc could be blocked by the protection strand. b) Once the protection strand is removed, the glue strand abc is active or on.
References
1. Jennifer E. Padilla, Matthew J. Patitz, Raul Pena, Robert T. Schwellery, Nadrian C. Seeman, Robert Sheline, Scott M. Summers, and Xingsi Zhong, Asynchronous Signal Passing for Tile Self-Assembly: Fuel Efficient Computation and Efficient Assembly of Shapes
2. J.E. Padilla, W. Liu, and N.C. Seeman, Hierarchical self assembly of patterns from the Robinson tilings: DNA tile design in an enhanced tile assembly model, Natural Computing online first, 17 August 2011 (2011).
3. G. Seelig, D. Soloveichik, D.Y. Zhang, and E. Winfree, Enzyme-free nucleic acid logic circuits, science 314 (2006), no. 5805, 1585.
4. P. Yin, H.M.T. Choi, C.R. Calvert, and N.A. Pierce, Programming biomolecular self-assembly path- ways, Nature 451 (2008), no. 7176, 318-322.
5. B. Yurke, A.J. Turberfield, A.P. Mills, F.C. Simmel, and J.L. Neumann, A DNA-fuelled molecular machine made of DNA, Nature 406 (2000), no. 6796, 605-608.
6. D.Y. Zhang and G. Seelig, Dynamic DNA nanotechnology using strand-displacement reactions, Nature Chemistry 3 (2011), no. 2, 103-113.
7. R.M. Dirks and N.A. Pierce, Triggered amplification by hybridization chain reaction, Proceedings of the National Academy of Sciences of the United States of America 101 (2004), no. 43, 15275.
8. G. Seelig, D. Soloveichik, D.Y. Zhang, and E. Winfree, Enzyme-free nucleic acid logic circuits, science 314 (2006), no. 5805, 1585.
9. L. Qian and E. Winfree, A simple DNA gate motif for synthesizing large-scale circuits, DNA Com- puting (2009), 70-89.
10. P. Yin, H.M.T. Choi, C.R. Calvert, and N.A. Pierce, Programming biomolecular self-assembly path- ways, Nature 451 (2008), no. 7176, 318-322.
11. K. Lund, A.J. Manzo, N. Dabby, N. Michelotti, A. Johnson-Buck, J. Nangreave, S. Taylor, R. Pei, M.N. Stojanovic, and N.G. Walter, Molecular robots guided by prescriptive landscapes, Nature 465 (2010), no. 7295, 206{210.
12. T. Omabegho, R. Sha, and N.C. Seeman, A bipedal DNA brownian motor with coordinated legs, Science 324 (2009), no. 5923, 67.
13. S.F.J. Wickham, M. Endo, Y. Katsuda, K. Hidaka, J. Bath, H. Sugiyama, and A.J. Turberfield, Di- rect observation of stepwise movement of a synthetic molecular transporter, Nature Nanotechnology 6 (2011), no. 3, 166-169.
14. P. Yin, H.M.T. Choi, C.R. Calvert, and N.A. Pierce, Programming biomolecular self-assembly path- ways, Nature 451 (2008), no. 7176, 318-322.